2022-05-04
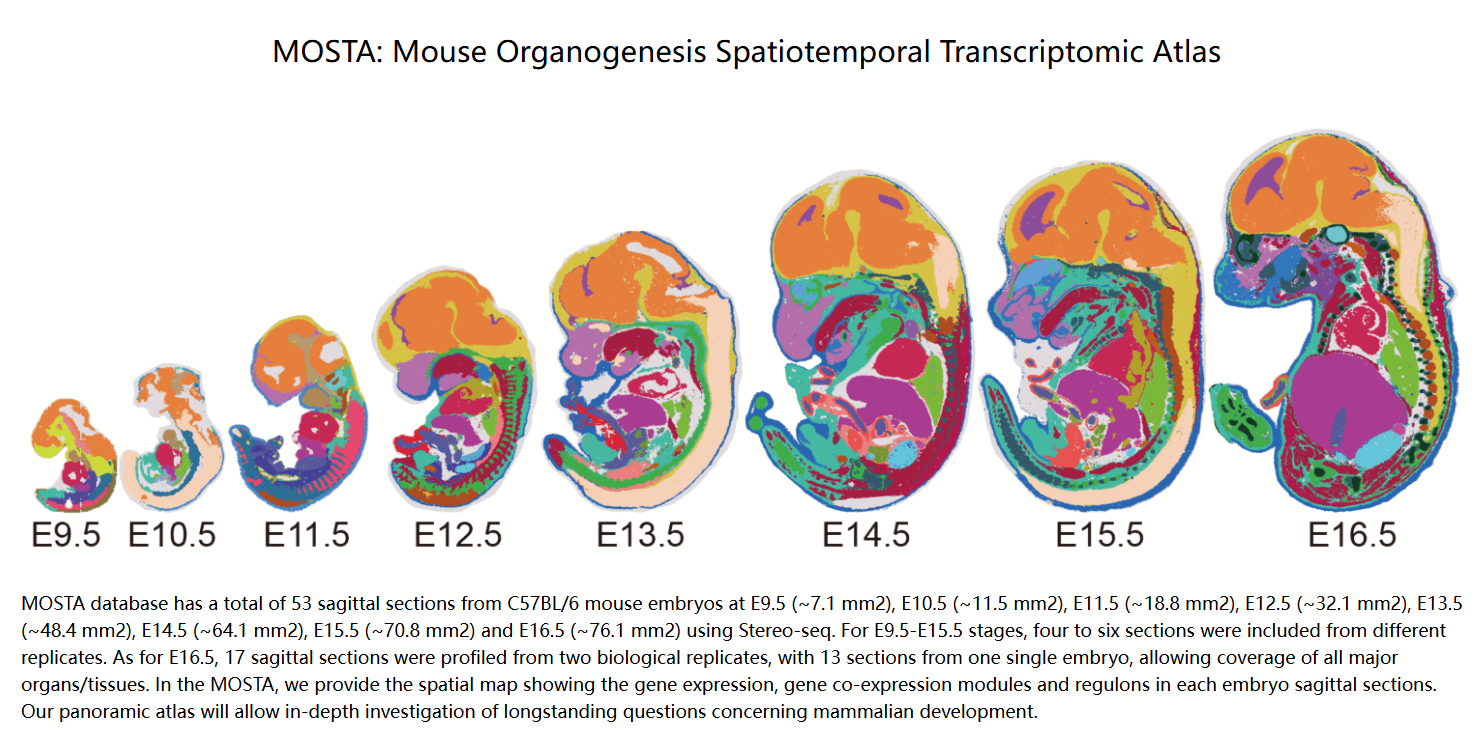
Chen, A., Liao, S., Cheng, M., Ma, K., Wu, L., Lai, Y., ... & Wang, J. (2022). Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell, 185(10), 1777-1792.
DOI:10.1016/j.cell.2022.04.003
2022-09-02
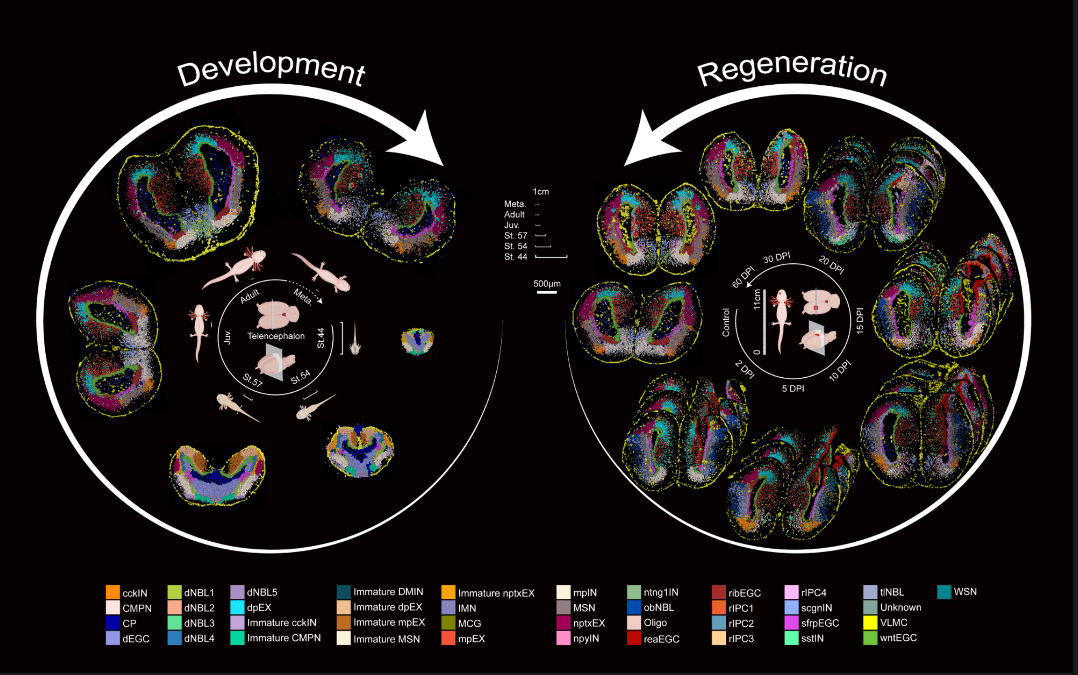
Wei, X., Fu, S., Li, H., Liu, Y., Wang, S., Feng, W., ... & Gu, Y. (2022). Single-cell Stereo-seq reveals induced progenitor cells involved in axolotl brain regeneration. Science, 377(6610), eabp9444.
DOI:10.1126/science.abp9444
2022-11-08
Lei, Y., Cheng, M., Li, Z., Zhuang, Z., Wu, L., Sun, Y., ... & Xu, X. (2022). Spatially resolved gene regulatory and disease-related vulnerability map of the adult Macaque cortex. Nature Communications, 13(1), 6747.
DOI:10.1038/s41467-022-34413-3
2022-05-04
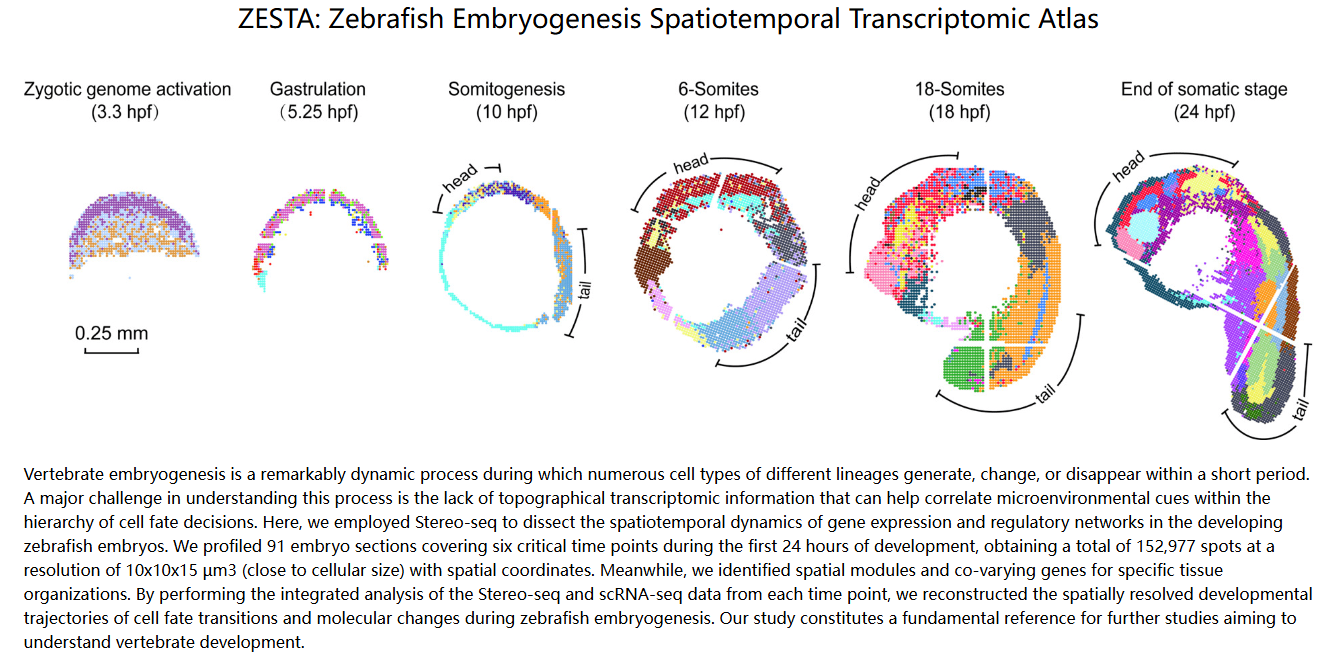
Liu, C., Li, R., Li, Y., Lin, X., Zhao, K., Liu, Q., ... & Liu, L. (2022). Spatiotemporal mapping of gene expression landscapes and developmental trajectories during zebrafish embryogenesis. Developmental Cell, 57(10), 1284-1298.
DOI:10.1016/j.devcel.2022.04.009
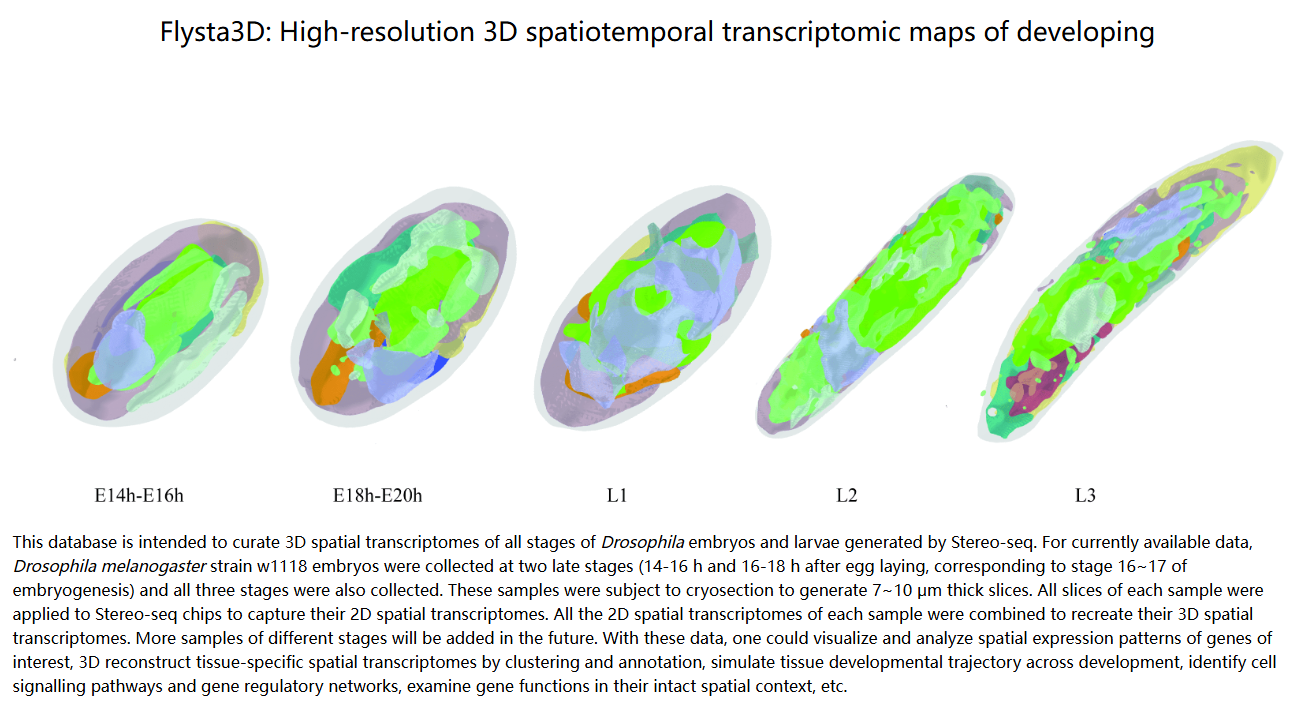
2022-05-04
Wang, M., Hu, Q., Lv, T., Wang, Y., Lan, Q., Xiang, R., ... & Liu, L. (2022). High-resolution 3D spatiotemporal transcriptomic maps of developing Drosophila embryos and larvae. Developmental Cell, 57(10), 1271-1283.
DOI:10.1016/j.devcel.2022.04.006
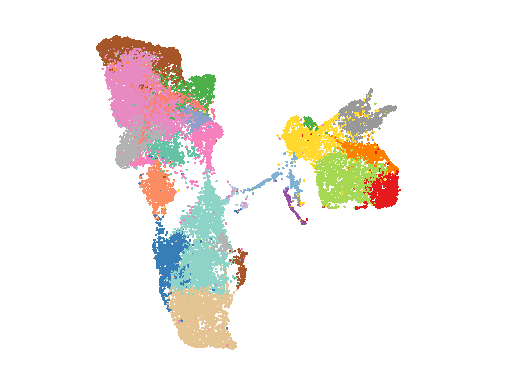
2022-01-14
Xia, Keke et al. “The single-cell stereo-seq reveals region-specific cell subtypes and transcriptome profiling in Arabidopsis leaves.” Developmental cell vol. 57,10 (2022): 1299-1310.e4.
DOI:10.1016/j.devcel.2022.04.011
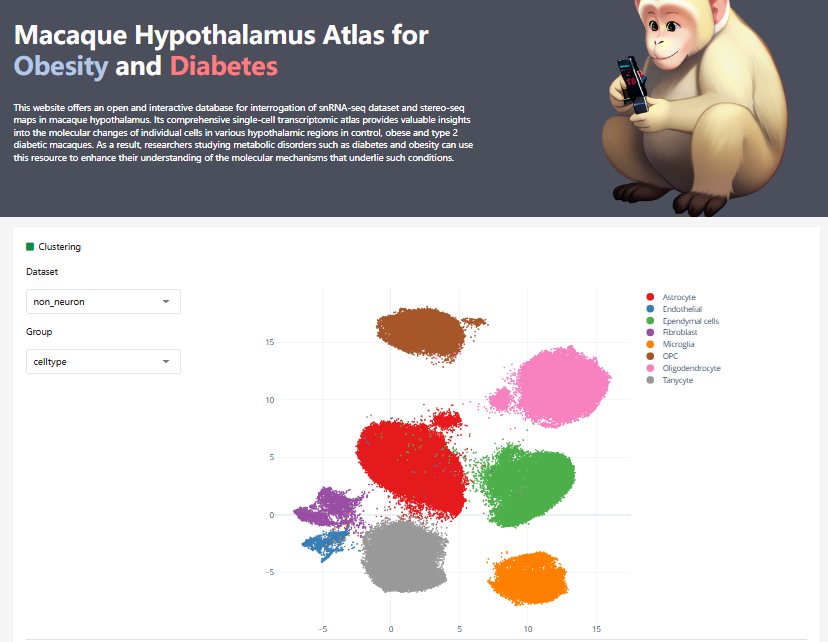
2024-02-06
Zhang, Xianglong et al. “A transcriptomic and proteomic atlas of obesity and type 2 diabetes in cynomolgus monkeys.” Cell reports vol. 42,8 (2023): 112952.
DOI:10.1016/j.celrep.2023.112952
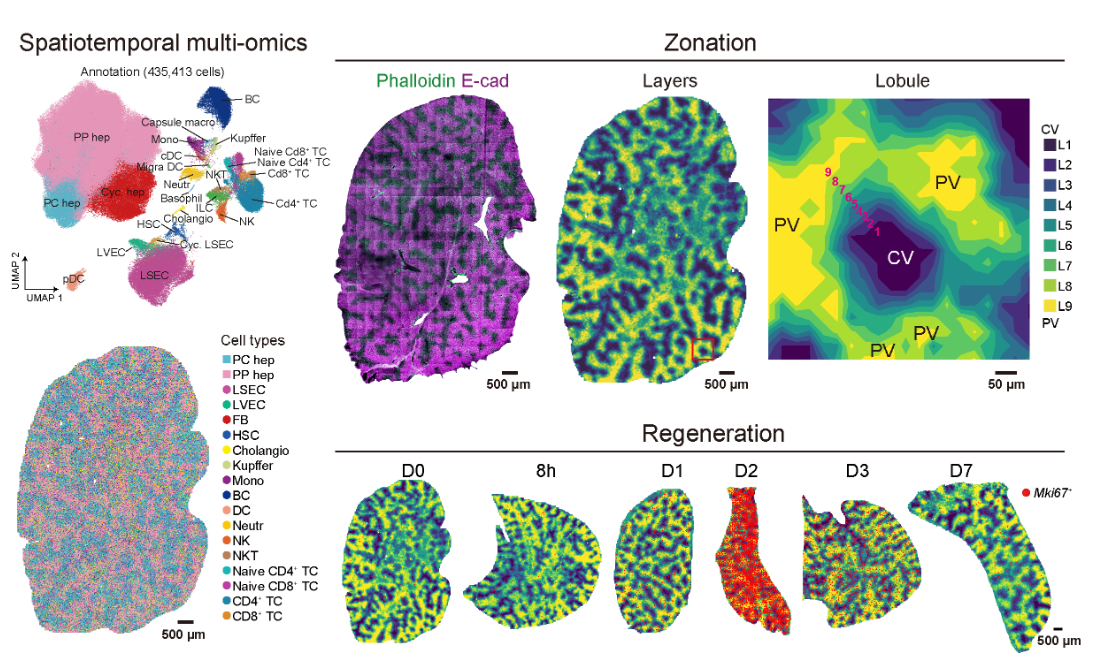
2024-04-16
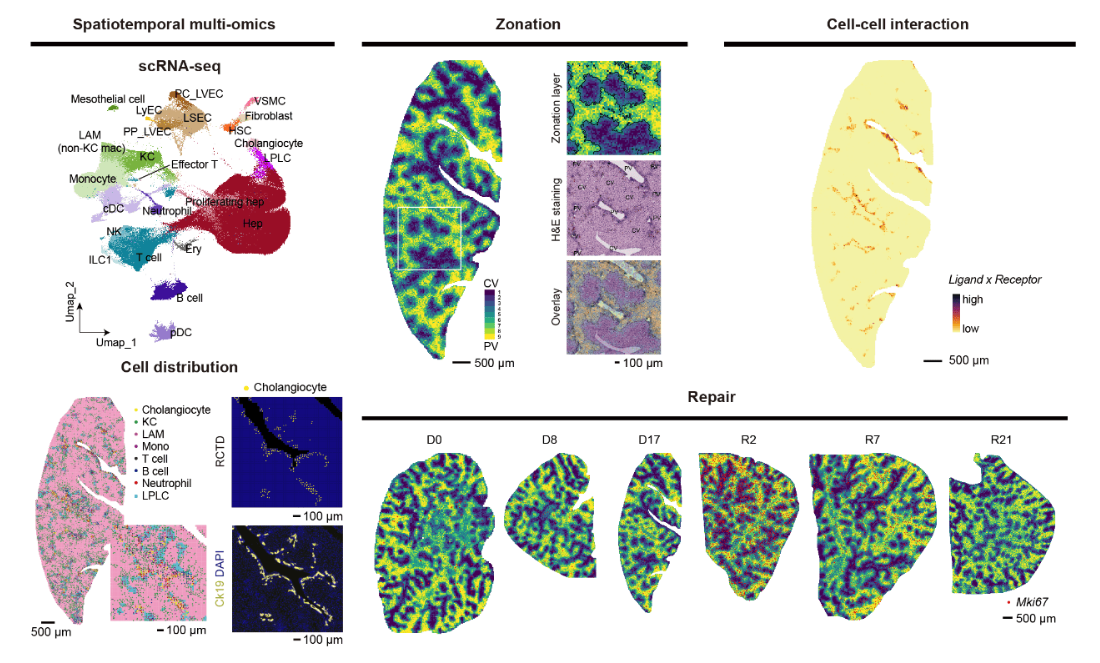
Xu, J., Guo, P., Hao, S. et al. A spatiotemporal atlas of mouse liver homeostasis and regeneration. Nat Genet (2024).
DOI:10.1038/s41588-024-01709-7
2024-04-16
Wu, B., Shentu, X., Nan, H. et al. A spatiotemporal atlas of cholestatic injury and repair in mice. Nat Genet (2024).
DOI:10.1038/s41588-024-01687-w
2024-05-14
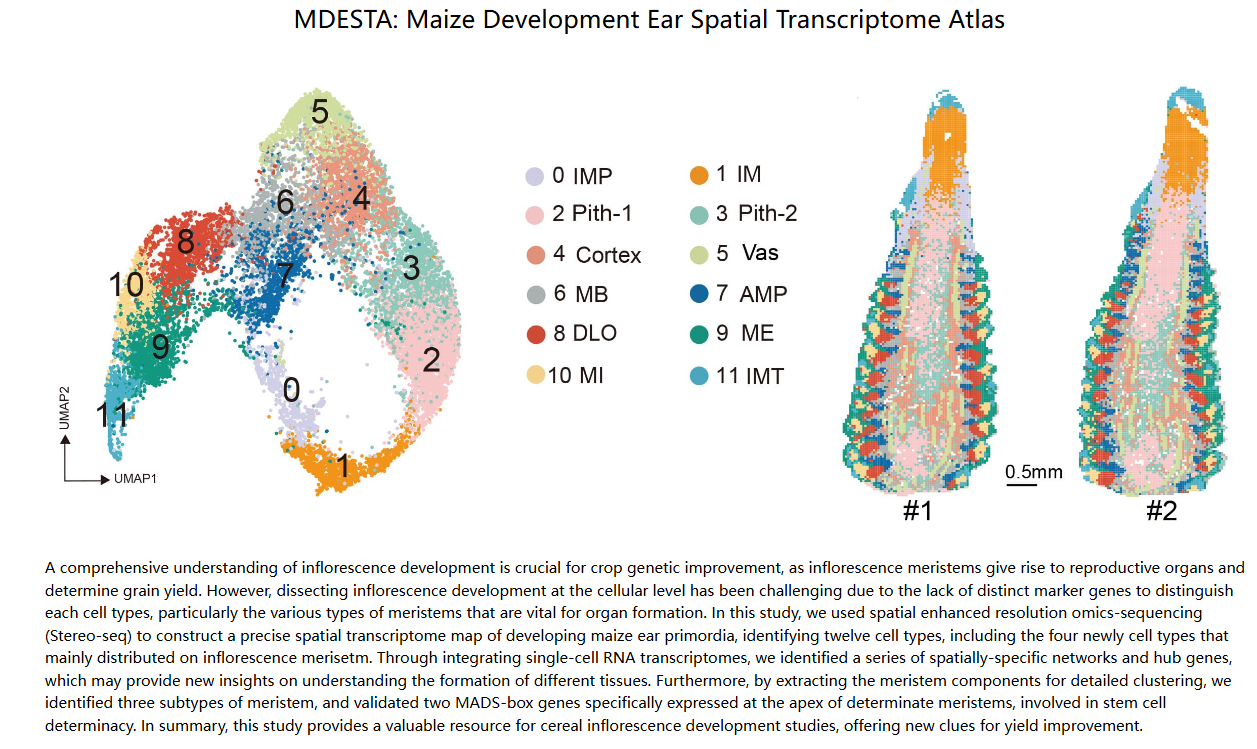
Wang, Y., Luo, Y., Guo, X. et al. A spatial transcriptome map of the developing maize ear. Nat. Plants (2024).
DOI:10.1038/s41477-024-01683-2

2024-09-27
Shijie Hao et al. ,Cross-species single-cell spatial transcriptomic atlases of the cerebellar cortex.Science385,eado3927(2024).
DOI:10.1126/science.ado3927
2024-10-22
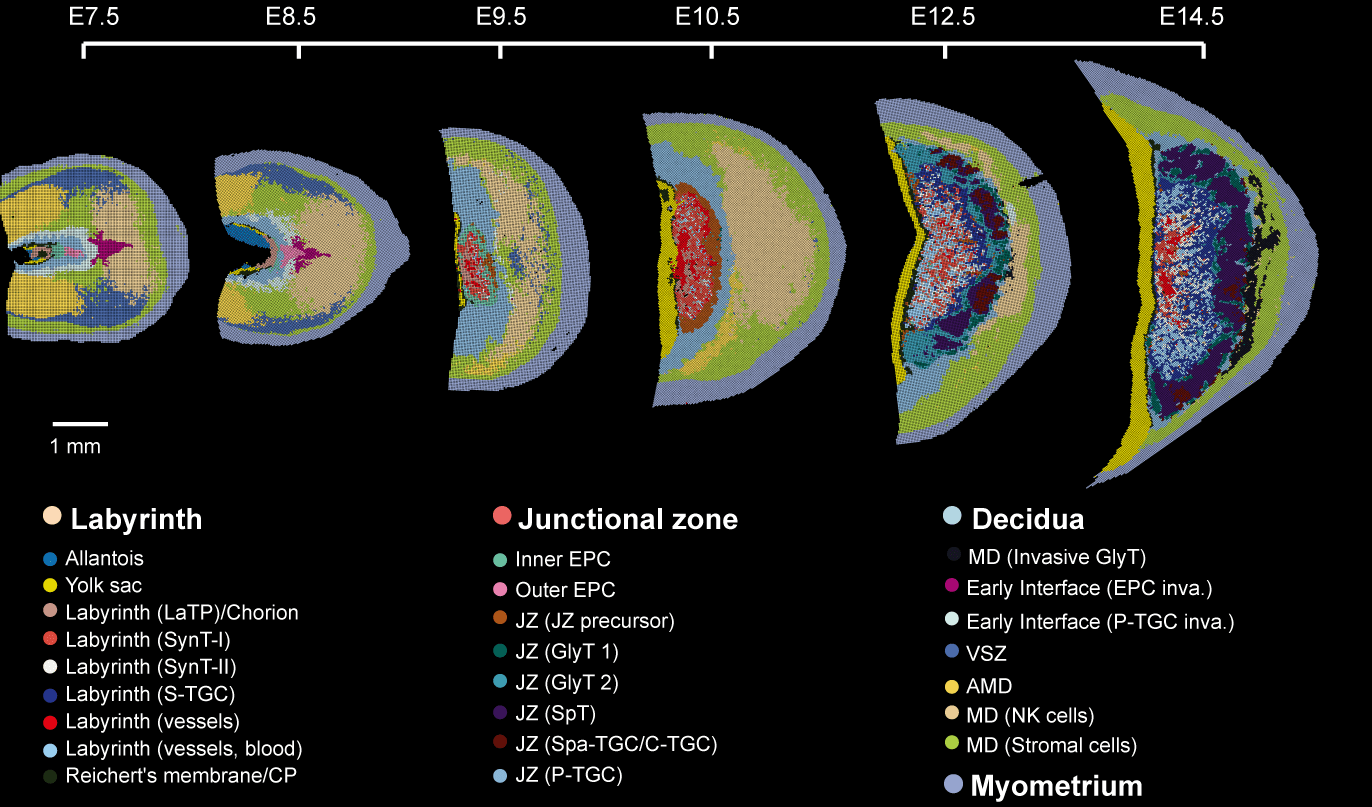
Wu, Y., Su, K., Zhang, Y. et al. A spatiotemporal transcriptomic atlas of mouse placentation. Cell Discov 10, 110 (2024).
DOI:10.1038/s41421-024-00740-6
2025-04-01
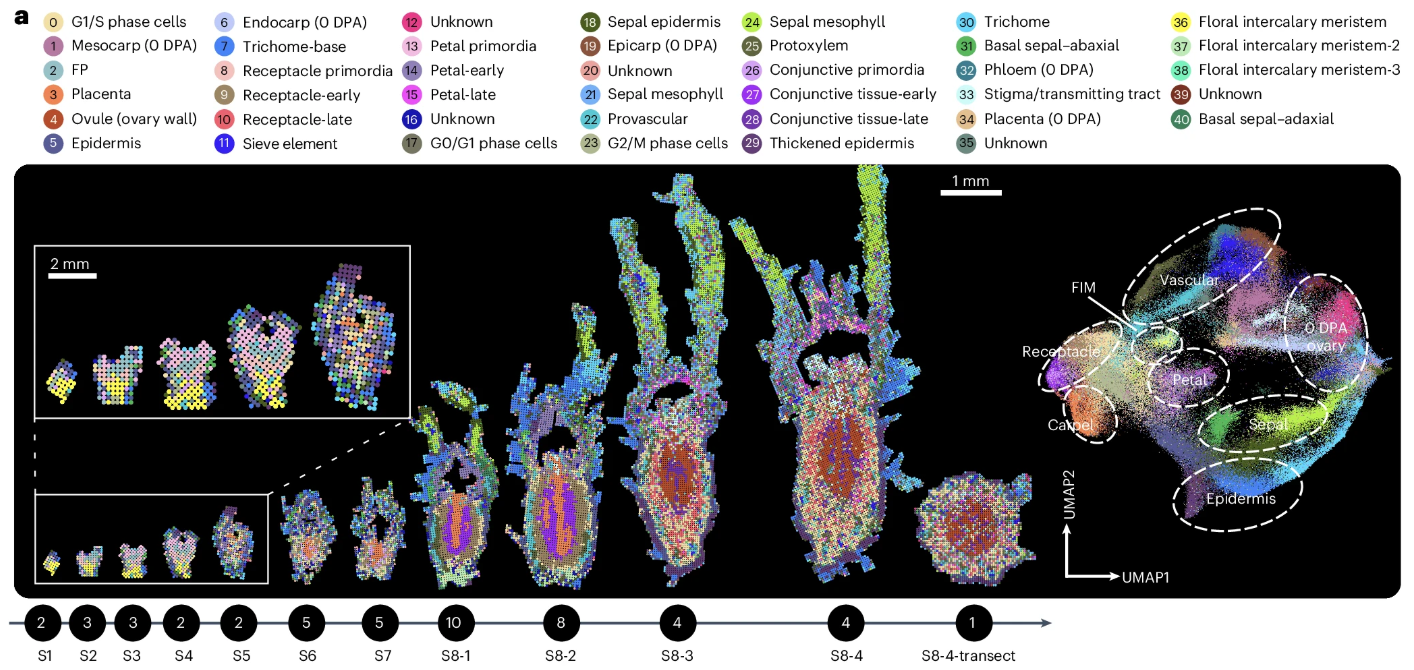
Dong, Z., Liu, X., Guo, X. et al. Developmental innovation of inferior ovaries and flower sex orchestrated by KNOX1 in cucurbits. Nat. Plants (2025).
DOI:10.1126/science.ado3927
2025-03-31
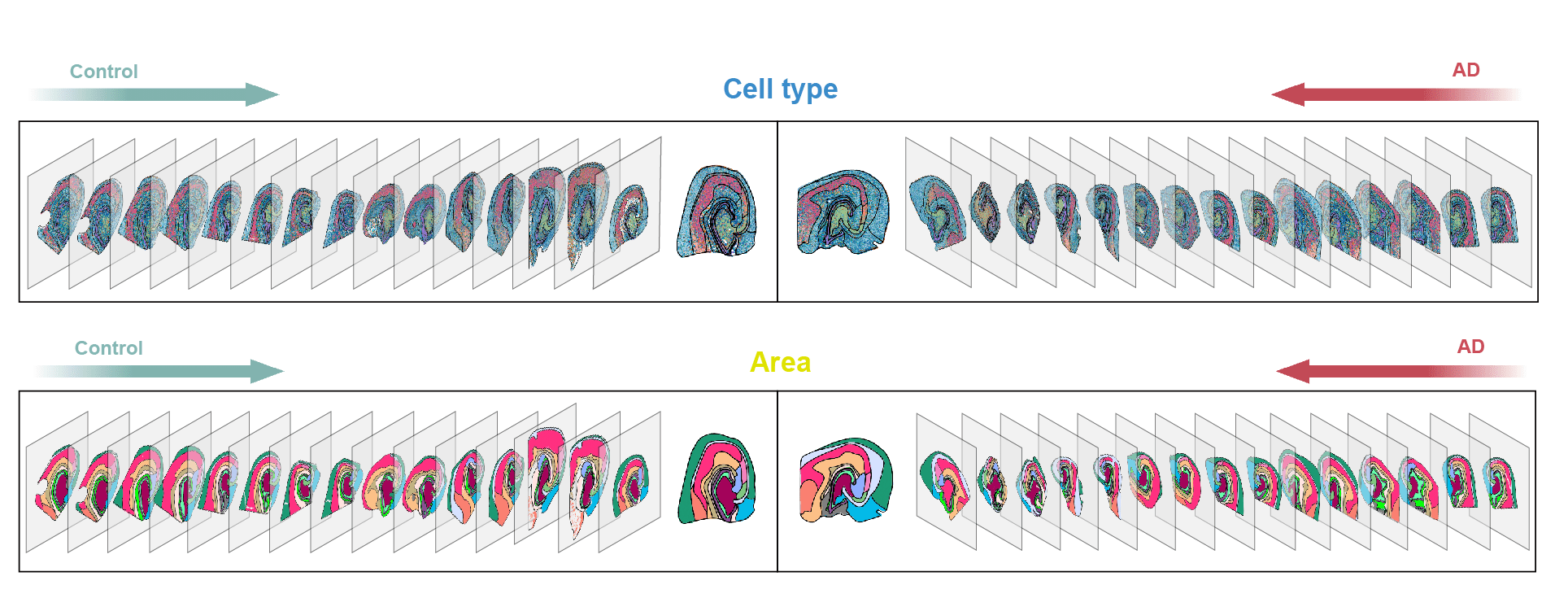
Wang, P., Han, L., Wang, L., Tao, Q., Guo, Z., Luo, T., ... & Zhang, J. (2025). Molecular Pathways and Diagnosis in Spatially Resolved Alzheimer’s Hippocampal Atlas. Neuron. Advance online publication.
DOI:10.1016/j.neuron.2025.03.002
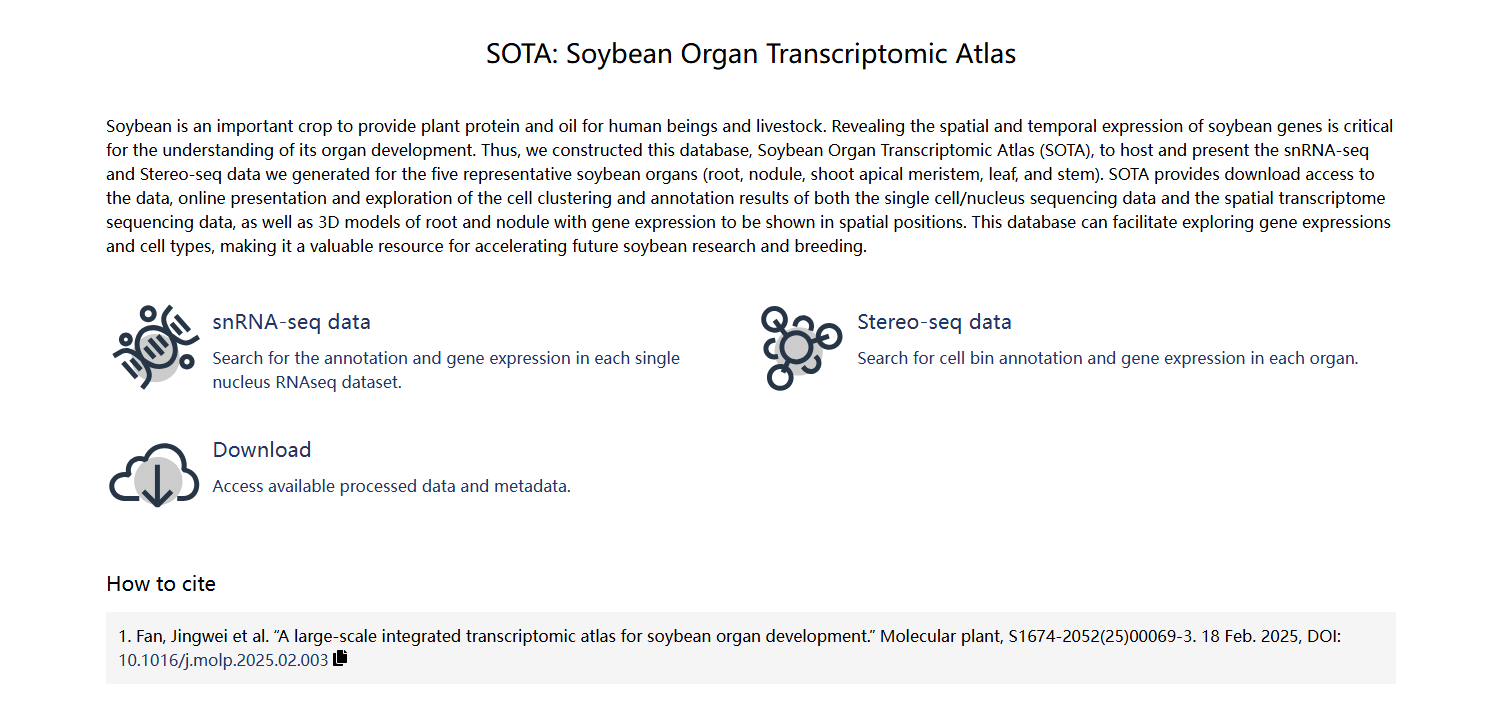

2025-02-18
1. Fan, Jingwei et al. “A large-scale integrated transcriptomic atlas for soybean organ development.” Molecular plant, S1674-2052(25)00069-3. 18 Feb.2025.
DOI:10.1016/j.molp.2025.02.003
2025-04-19
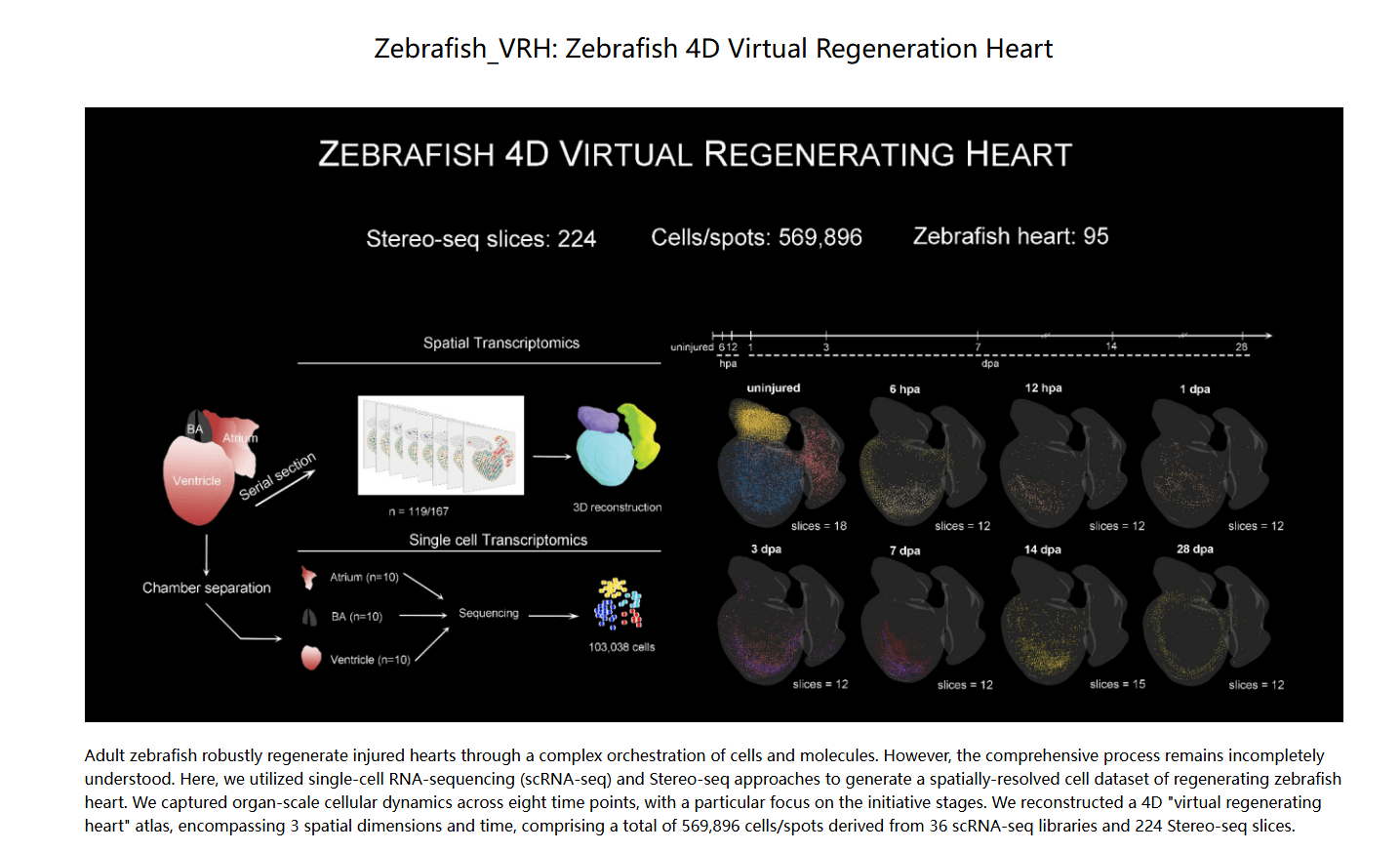
Li, L., Lu, M., Guo, L. et al. An organ-wide spatiotemporal transcriptomic and cellular atlas of the regenerating zebrafish heart. Nat Commun 16, 3716 (2025).
DOI:10.1038/s41467-025-59070-0
2025-07-07
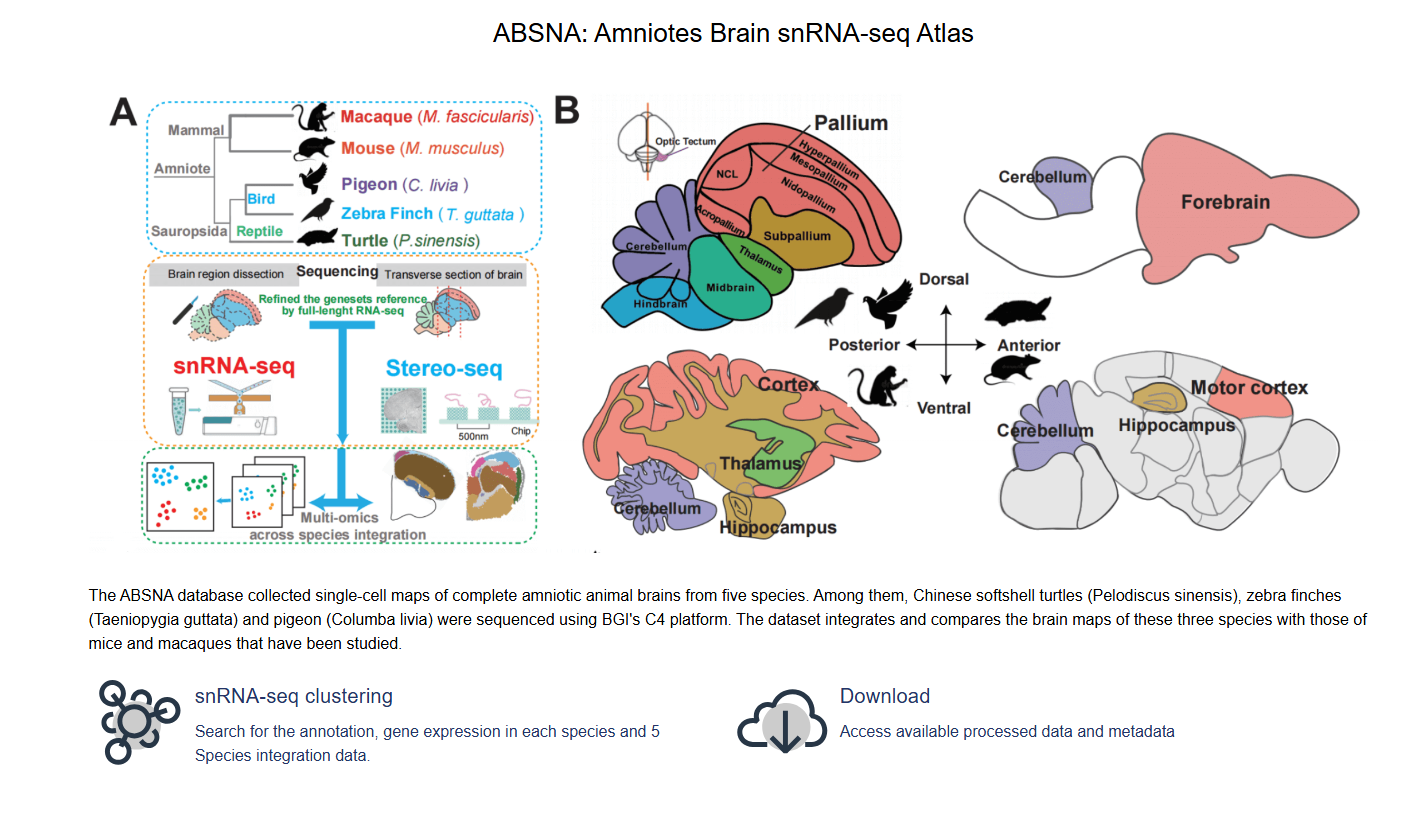
Chen D, Zhuang Z, Huang M, Huang Y, Yan Y, Zhang Y, Lin Y, Jin X, Wang Y, Huang J, Xu W, Pan J, Wang H, Huang F, Liao K, Cheng M, Zhu Z, Bai Y, Niu Z, Zhang Z, Xiang Y, Wei X, Yang T, Zeng T, Dong Y, Lei Y, Sun Y, Wang J, Yang H, Sun Y, Cao G, Poo M, Liu L, Naumann RK, Xu C, Wang Z, Xu X, Liu S. Genomic evolution reshapes cell-type diversification in the amniote brain. Dev Cell. 2025 Jul 7;60(13):1900-1915.e5.
DOI:10.1016/j.devcel.2025.04.014
2025-08-21
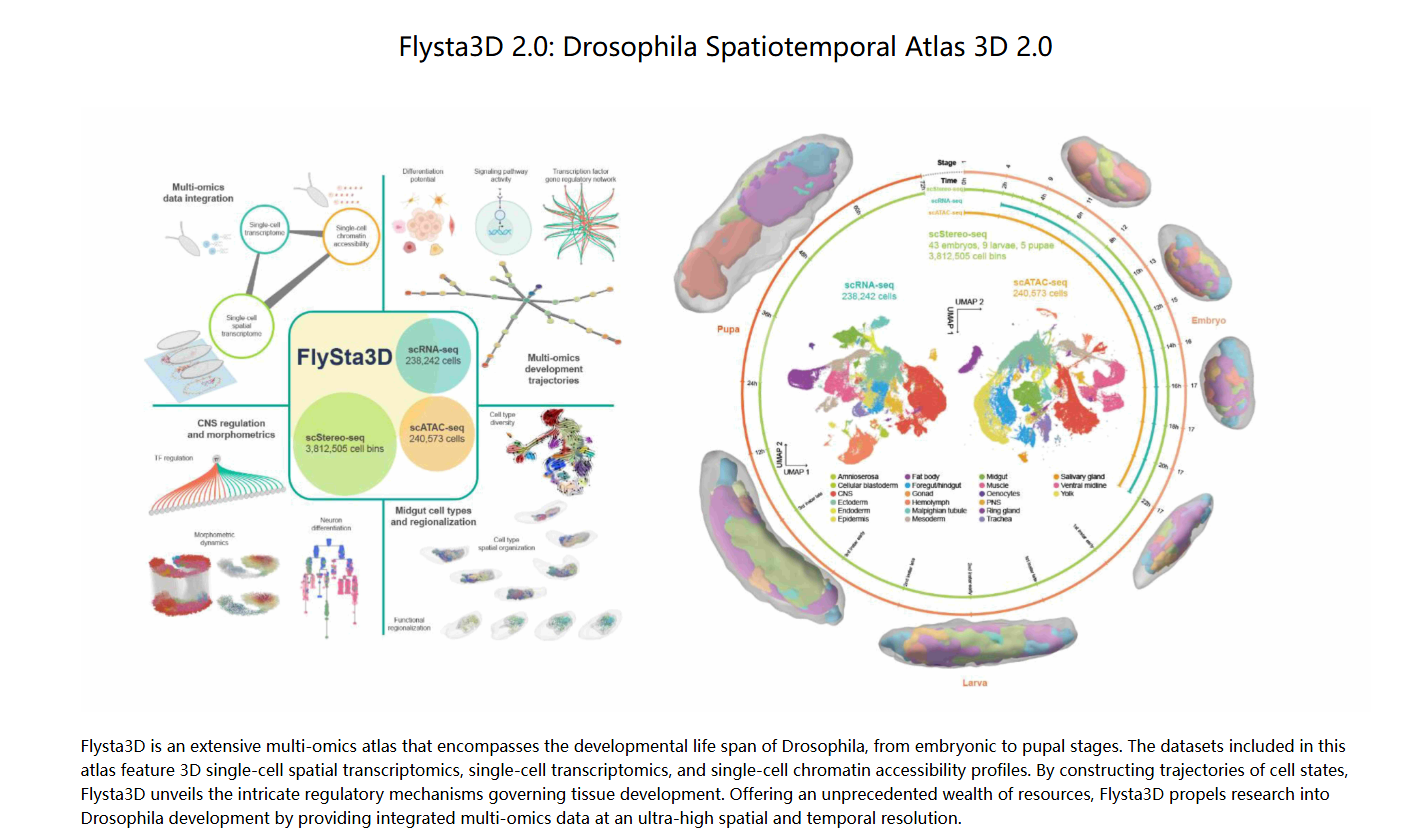
Wang M, Hu Q, Tu Z, Kong L, Yu T, Jia Z, Wang Y, Yao J, Xiang R, Chen Z, Zhao Y, Zhou Y, Ye Q, Ouyang K, Wang X, Bai Y, Yang Z, Wang H, Wang Y, Jiang H, Yang T, Chen J, Huang Y, Yin N, Mo W, Liang W, Liu C, Lin X, Liu C, Gu Y, Chen W, Liu L, Xu X, Hu Y. A Drosophila single-cell 3D spatiotemporal multi-omics atlas unveils panoramic key regulators of cell-type differentiation. Cell. 2025 Aug 21;188(17):4734-4753.e31.
DOI::10.1016/j.cell.2025.05.04
2025-08-21
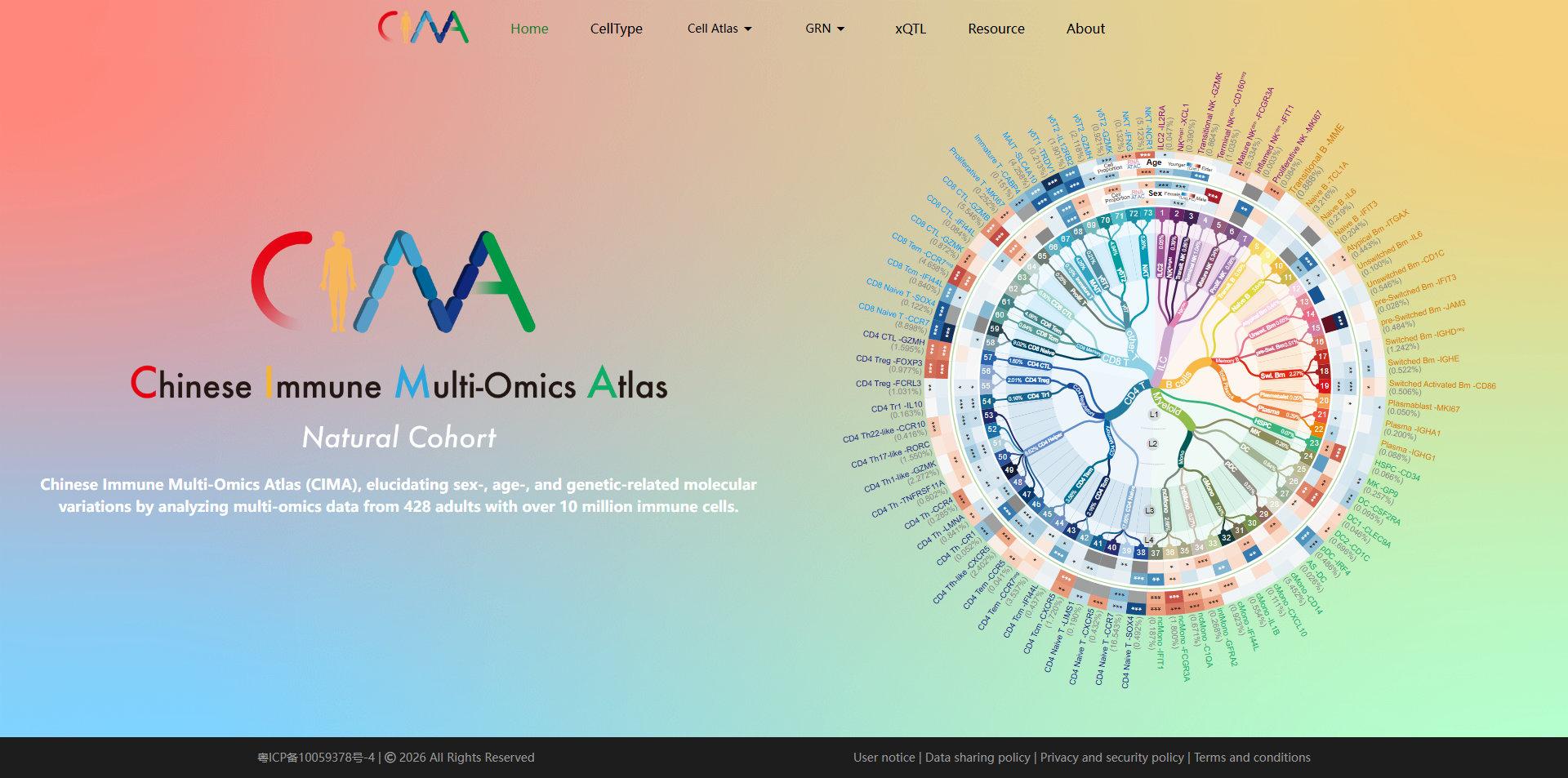
Yin J, Zheng Y, Huang Z, Zhou W, Yuan Y, Cai P, Bai Y, Yang S, Gao Y, Duan S, Wang Y, Xu Z, Zhang W, Zhang X, Wei Y, Huang Y, Liu Y, Wang W, Yang T, Zhang Z, Chen X, Zhang X, Lv J, Li F, Zhang Y, Zeng G, Wang X, Ma W, Hou G, Hao S, Liu C, Lai Y, Liu P, Wang B, Li Y, Zhang W, Gao P, Xie J, Esteban MA, Gu Y, Liu X, Ji J, Qi T, Liu B, Wang H, Zhao Y, Yang X, Wang X, Chen R, Yang J, Yin Y, Wang J, Cao Y, Xu X, Liu L, Jin X, Liu C. Chinese Immune Multi-Omics Atlas. Science. 2026 Jan 8;391(6781):eadt3130.
DOI:10.1126/science.adt3130
2026-04-16
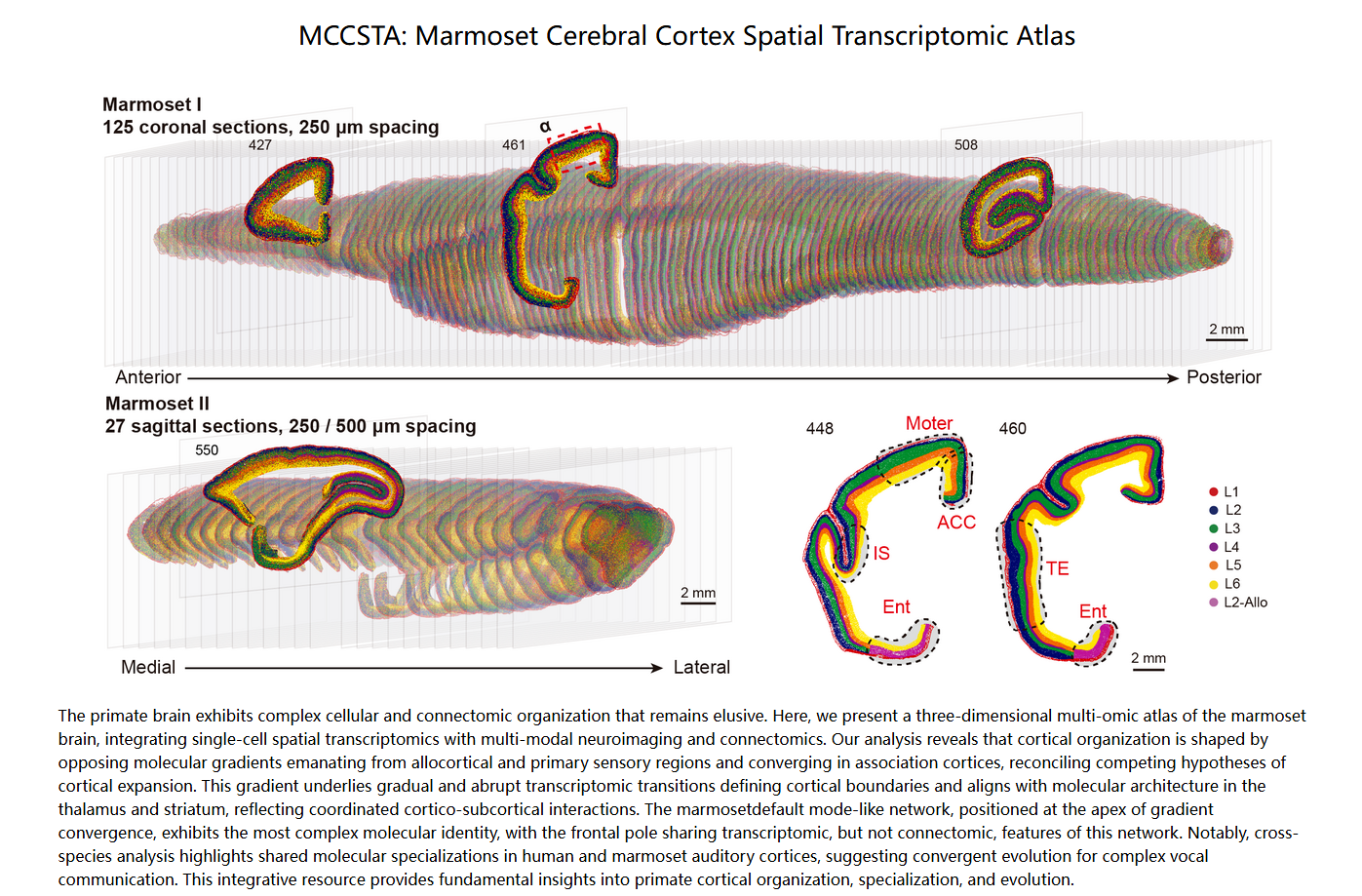
Zhi Huang et al. ,An opposing molecular gradient axis underlies primate cortical organization.Science392,eaea2673(2026).
DOI:10.1126/science.aea2673
