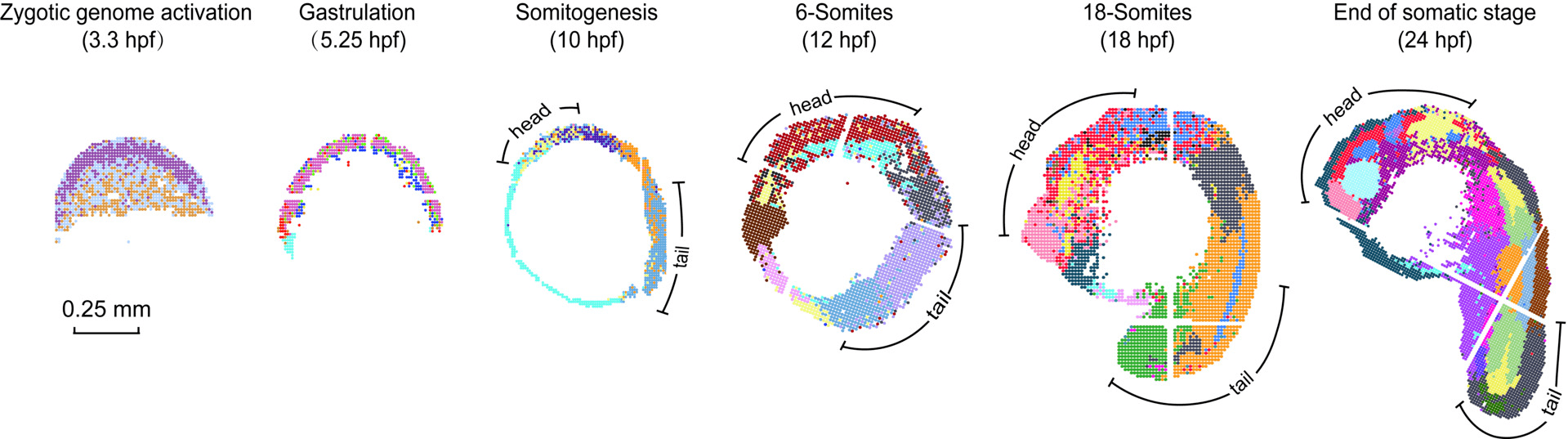
Vertebrate embryogenesis is a remarkably dynamic process during which numerous cell types of different lineages generate, change, or disappear within a short period. A major challenge in understanding this process is the lack of topographical transcriptomic information that can help correlate microenvironmental cues within the hierarchy of cell fate decisions. Here, we employed Stereo-seq to dissect the spatiotemporal dynamics of gene expression and regulatory networks in the developing zebrafish embryos. We profiled 91 embryo sections covering six critical time points during the first 24 hours of development, obtaining a total of 152,977 spots at a resolution of 10x10x15 µm3 (close to cellular size) with spatial coordinates. Meanwhile, we identified spatial modules and co-varying genes for specific tissue organizations. By performing the integrated analysis of the Stereo-seq and scRNA-seq data from each time point, we reconstructed the spatially resolved developmental trajectories of cell fate transitions and molecular changes during zebrafish embryogenesis. Our study constitutes a fundamental reference for further studies aiming to understand vertebrate development.

Search for the annotation and gene expression on each zebrafish embryo covering six critical time points during the first 24 hours of development in Stereo-seq and scRNA-seq data.

Protocol of the Stereo-seq chip generation and library preparation.

We have made our raw data and software resource available for analyse the raw data for the research community.

We provided our processed data and meta data available for the research community.
How to cite
