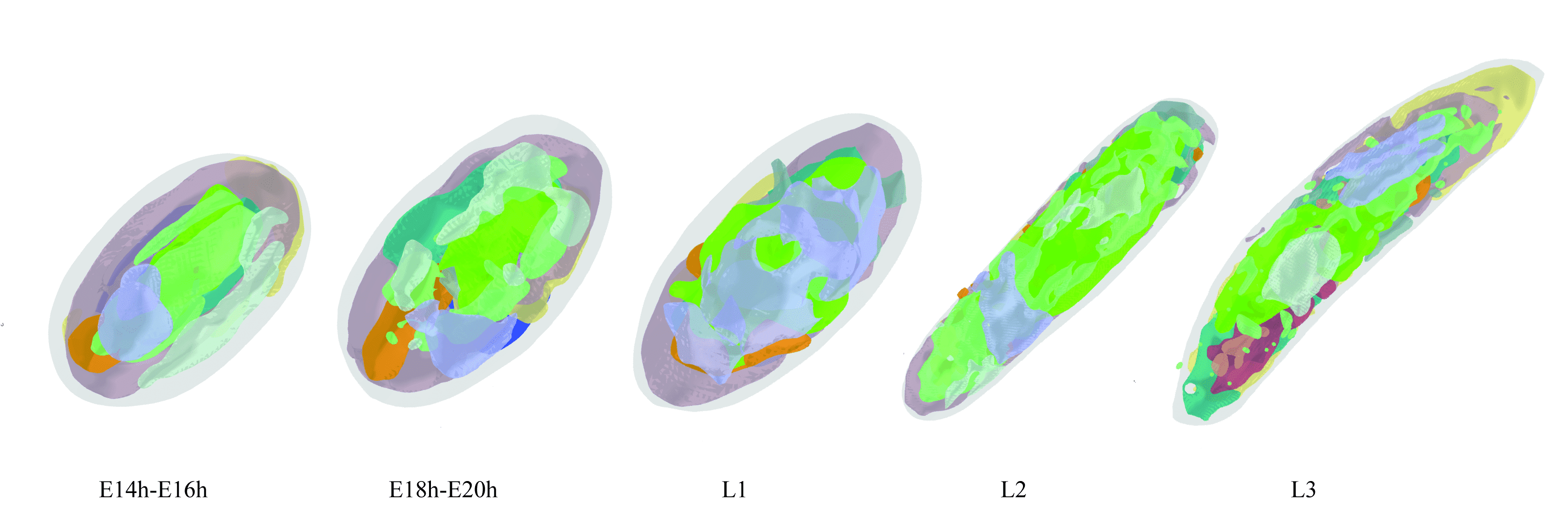
This database is intended to curate 3D spatial transcriptomes of all stages of Drosophila embryos and larvae generated by Stereo-seq. For currently available data, Drosophila melanogaster strain w1118 embryos were collected at two late stages (14-16 h and 16-18 h after egg laying, corresponding to stage 16~17 of embryogenesis) and all three stages were also collected. These samples were subject to cryosection to generate 7~10 μm thick slices. All slices of each sample were applied to Stereo-seq chips to capture their 2D spatial transcriptomes. All the 2D spatial transcriptomes of each sample were combined to recreate their 3D spatial transcriptomes. More samples of different stages will be added in the future. With these data, one could visualize and analyze spatial expression patterns of genes of interest, 3D reconstruct tissue-specific spatial transcriptomes by clustering and annotation, simulate tissue developmental trajectory across development, identify cell signalling pathways and gene regulatory networks, examine gene functions in their intact spatial context, etc.

Search for tissue annotation and gene expression at each developmental stage.

Protocols for Stereo-seq chip generation and library preparation.

Visualize 3D reconstructed models. Access available raw data and data analysis softwares.

Access available processed data and metadata
How to cite
