About NHPCA
Non-human primate cell atlas (NHPCA) is a single cell transcriptomics data resource that provides visualization and preliminary analysis of transcriptomic and forthcoming epigenetic single-cell data sampled from NHP organs or tissues, aiming to construct a comprehensive, high quality single cell reference map of the NHP body. All raw data have been deposited to CNGB Nucleotide Sequence Archive (accession code: CNP0001469).
Data
We have generated the largest single-cell/single-nucleus transcriptomic NHPCA dataset (Macaca fascicularis) so far, encompassing 1,144,706 cells/nuclei that passed a QC cutoff of 500 genes and 10% mitochondrial genes from 45 organs/tissues belonging to all major systems (nervous, immune, endocrine, cardiovascular, respiratory, digestive, skeletal, reproductive and urinary). All these single cell data are welcome to download or upload for analysis and comparison.
Data exploration
To facilitate the exploration of this dataset, we have created an open and interactive single cell database that will allow everybody to explore gene expression and species comparison in single-cell level. We have illustrated either global or individual clustering and annotation from this dataset. Species comparison of gene expression and cell-cell interactions are also included in this database. Based on this, researchers can explore gene expression profiles of their interested cell types and compare their own single cell data with monkey through the upload data function.
How to cite
Han, L., Wei, X., Liu, C. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature (2022). https://doi.org/10.1038/s41586-022-04587-3
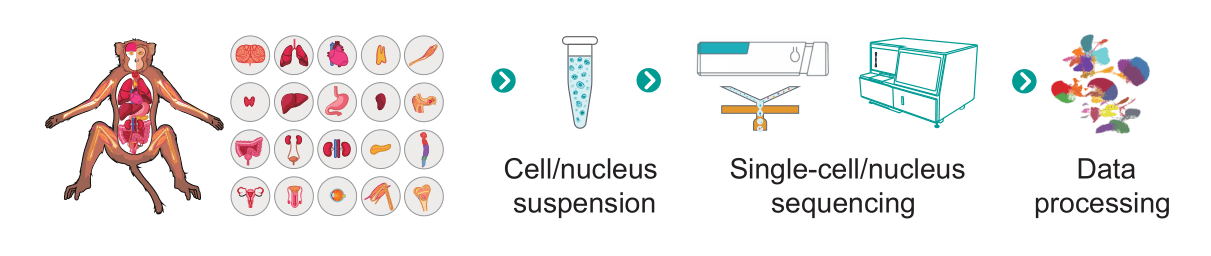 Workflow of sn/scRNA-seq
Workflow of sn/scRNA-seq