The MMHP was initiated by Professor Mathias Uhlen and Professor Lars Engstrand from the Karolinska Institute of Sweden in partnership with BGI, and scientists from Denmark, France, Latvia and China. The MMHP was officially launched during the ICG-14 meeting (October, 2019) by Professor Lars Engstrand and Professor Guang Ning, where the MMHP founding members agreed to release sequencing data of 10,000 meta genomes comprising samples from Sweden, Denmark, Latvia, and China among MMHP members to initiate the project.
An increasing number of studies has highlighted the important role of the microbiome in human health and disease. However, information on the composition of the microbiome in different parts of the body across countries, ages, sexes, and in relation to human health and disease is still limited. More importantly, though thousands of peer-reviewed papers have been published, very few open access databases are available. With the featured MGI DNB™ sequencing technology, the MMHP aims to sequence and analyze one million microbiome samples in the coming 3-5 years, focusing on feces and saliva (other body sites for which large number of samples can be obtained will be accepted), and make the first comprehensive map of the human microbiome publicly available.
The MMHP envisions to comprise members from academic and private institutions around the world. It is only by working together that we will be able to achieve the ambitious goals of this project. The MMHP encourages members to share microbiome sequence data through the data coordination platform across MMHP working groups and make data publicly available in regular releases. The MMHP will have an annual meeting to review the progress of the project and plan the next phases of the project.
The MMHP is steered and governed by an Organizing Committee (OC), which is the decision-making body of the MMP.
The Scientific Committee (SC) organizes samples, clinical phenotypes, working groups, and data analysis teams.
The Executive Committee (EC) organizes meetings, sequencing resources, recommends experimental designs, analysis frameworks and best practices for different aspects and needs of the MMHP, from data collection to the data coordination platform.
The project will rely mainly on MGI's DNBSEQ™ microbial genome sequencing technology to construct human microbial maps of different populations and health conditions. These microbial maps make it possible to establish a baseline of microecology research at the large-scale population level in order to promote research of cutting-edge translational medicine in the field of the human microbiome.
Comparisons between Hiseq and DNBSEQ™ have been done by researchers. The high accuracy and technical reproducibility shown by Fang C, et al confirm the applicability of the DNBSEQ™ for metagenomic studies1. Patterson J, et al have showcased DNBSEQ™ as a viable alternative to HiSeq for de novo RNA-Seq assembly2. Both work have shown that DNBSEQ™ gives better performance in high GC regions.
In addition, MGI offers convenient Kit for sample collection and preservation in room temperature for 14 days3.
For more information, please visit the MGI homepage (https://www.mgitech.cn/).
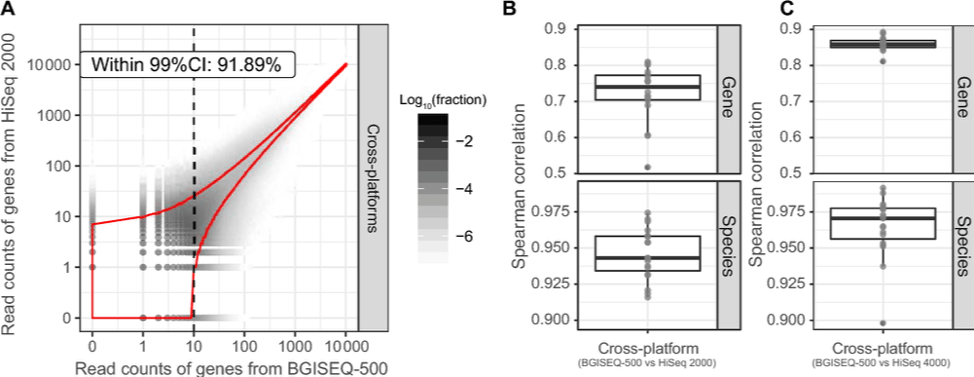
Fang C, et al. 2018
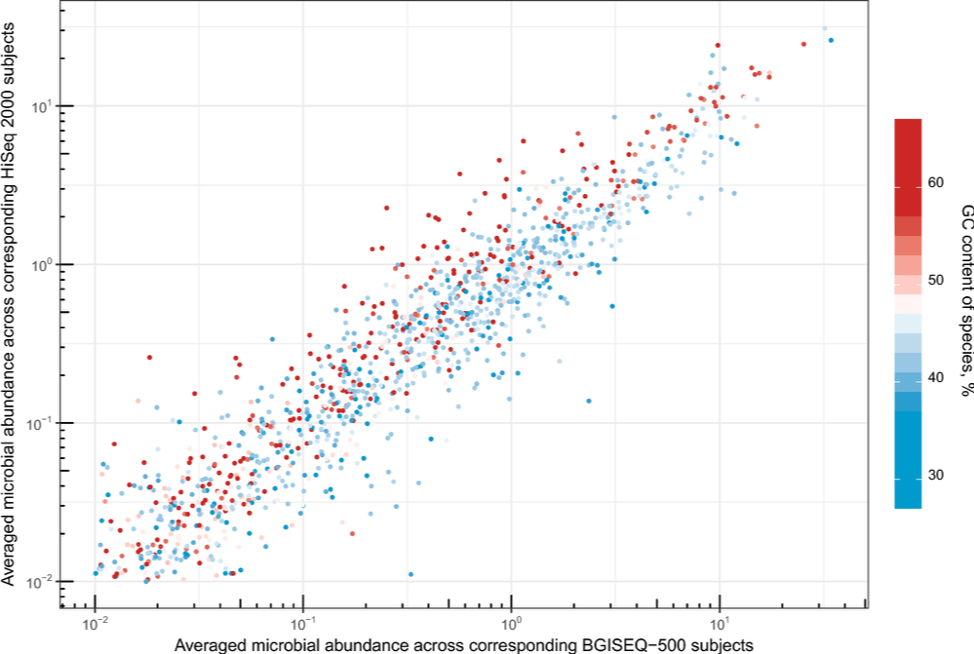
Fang C, et al. 2018
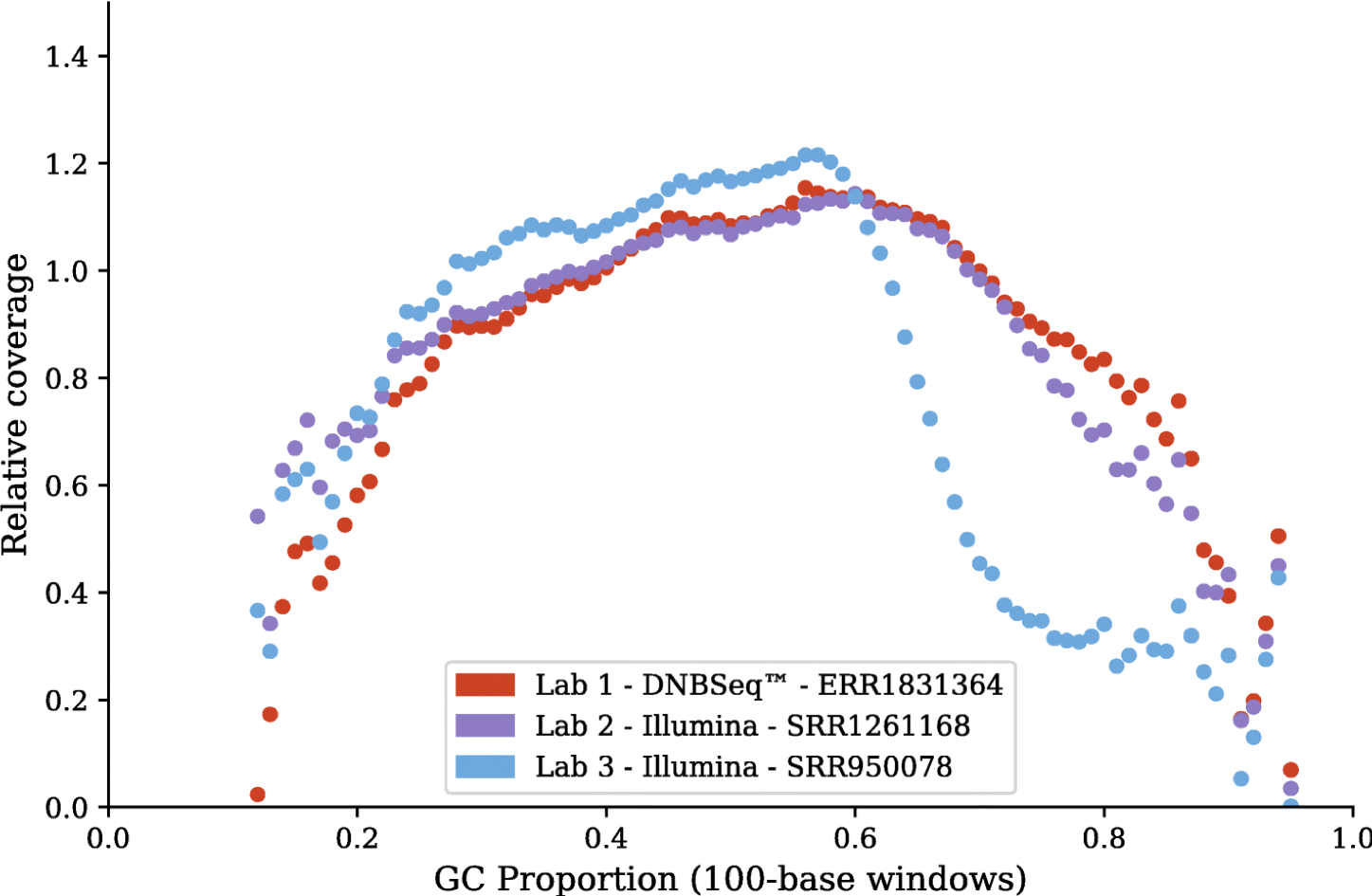
Patterson J, et al.
Reference
- Fang C, Zhong H, Lin Y, et al. Assessment of the cPAS-based BGISEQ-500 platform for metagenomic sequencing. Gigascience, 2018, 7(3): gix133.
- Patterson J, Carpenter E J, Zhu Z, et al. Impact of sequencing depth and technology on de novo RNA-Seq assembly. BMC genomics, 2019, 20(1): 604.
- Han M, Hao L, Lin Y, et al. A novel affordable reagent for room temperature storage and transport of fecal samples for metagenomic analyses. Microbiome, 2018, 6(1): 43.