help
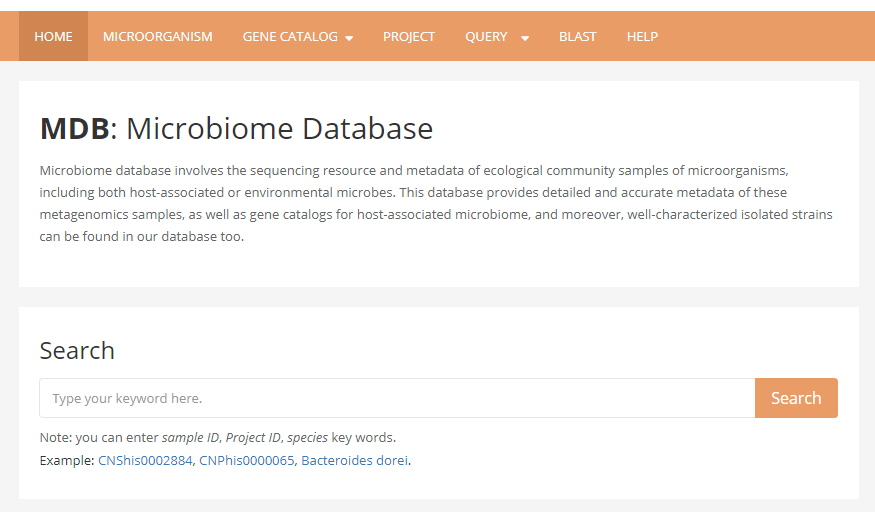
1 Home page contains a brief introduction of Microbiome Database website, as well as a search box. User can type keywords in it, including sample ID, project ID or microorganism name, information of matched samples or microorganisms will be listed as results.
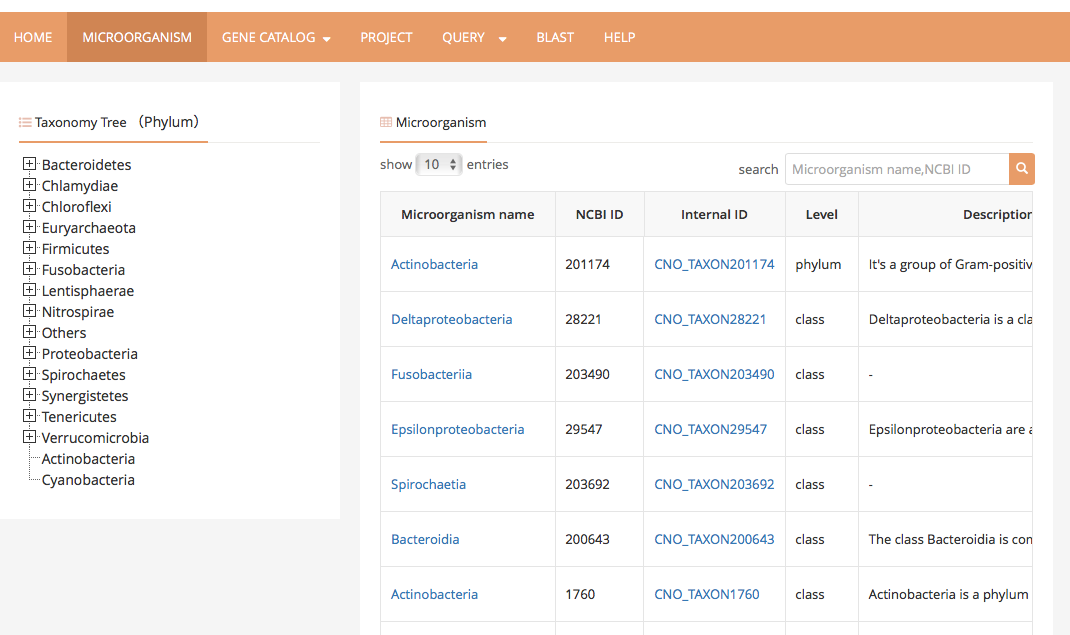
2 Microorganism page contains a taxonomy tree as a navigator bar on the left, and a search box with result area on the right. User can choose targeted microorganisms by unfolding taxonomy tree or typing name in search box. Hit microorganisms will be listed at the result area.
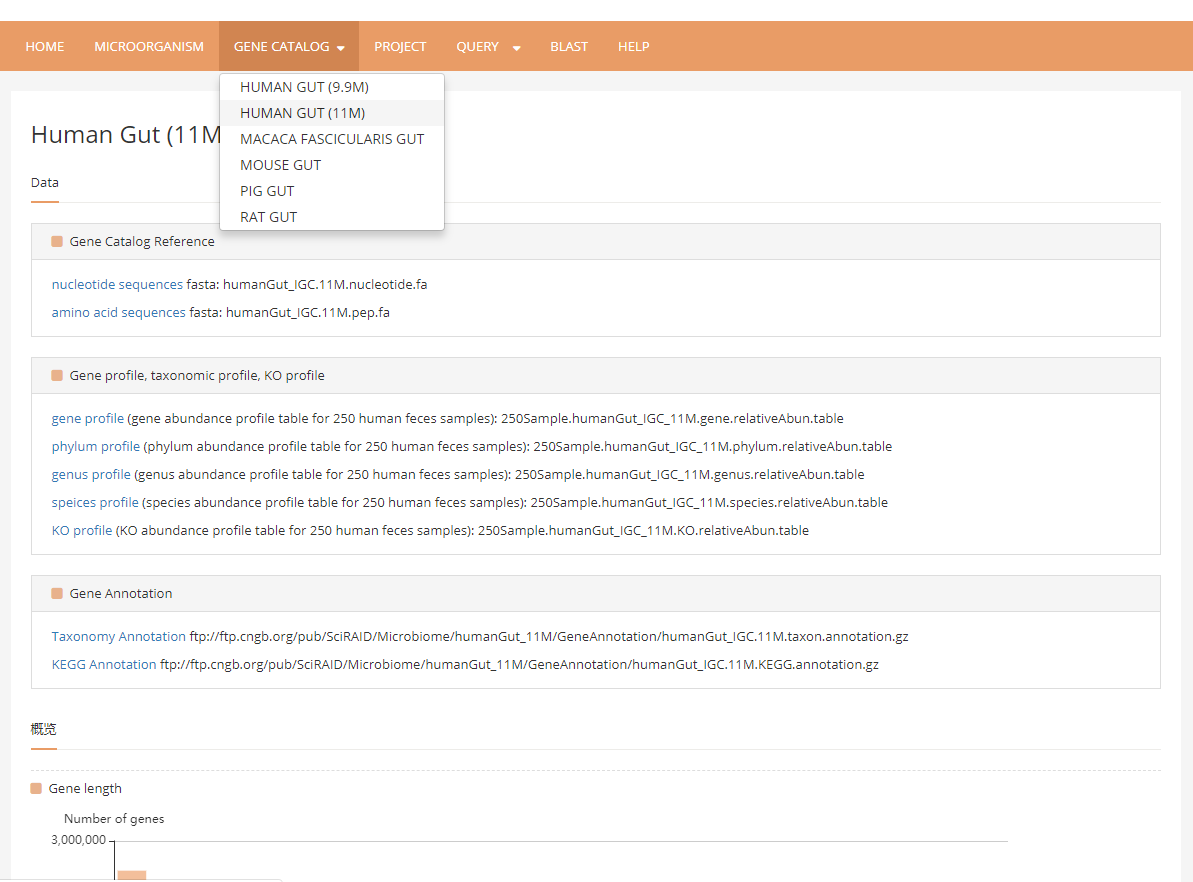
3 Gene catalog page contains several microbiome gene catalogs assembled from metagenomics data of human and model-animals, including FASTA files, annotation and abundance profiles.
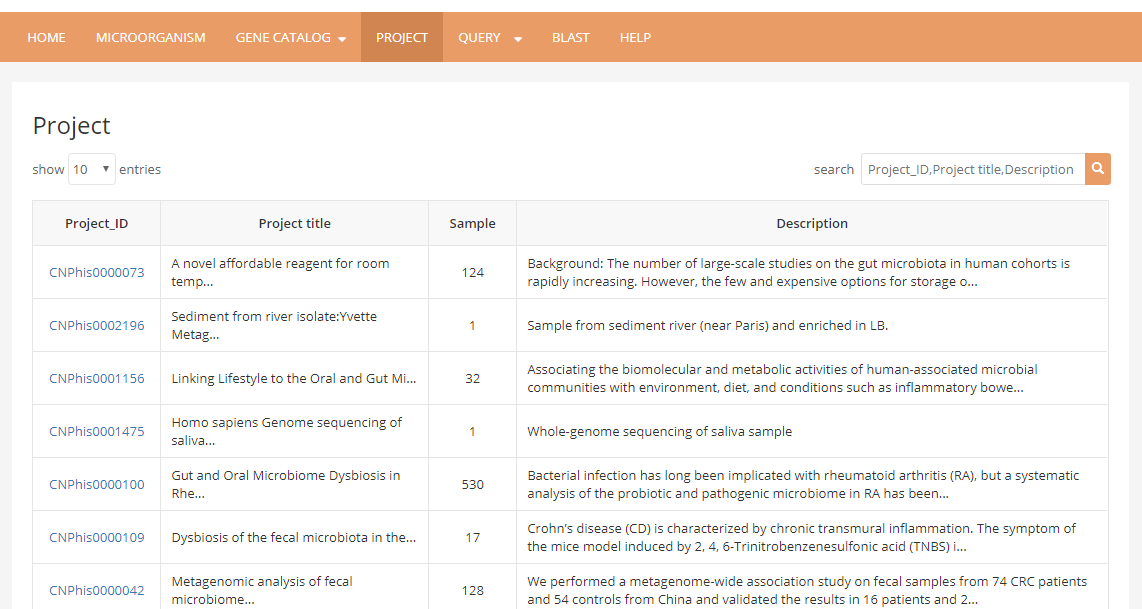
4 User can use Project page to filter projects in Microbiome Database by project name at this moment, all the samples belong to this certain project will be listed below.
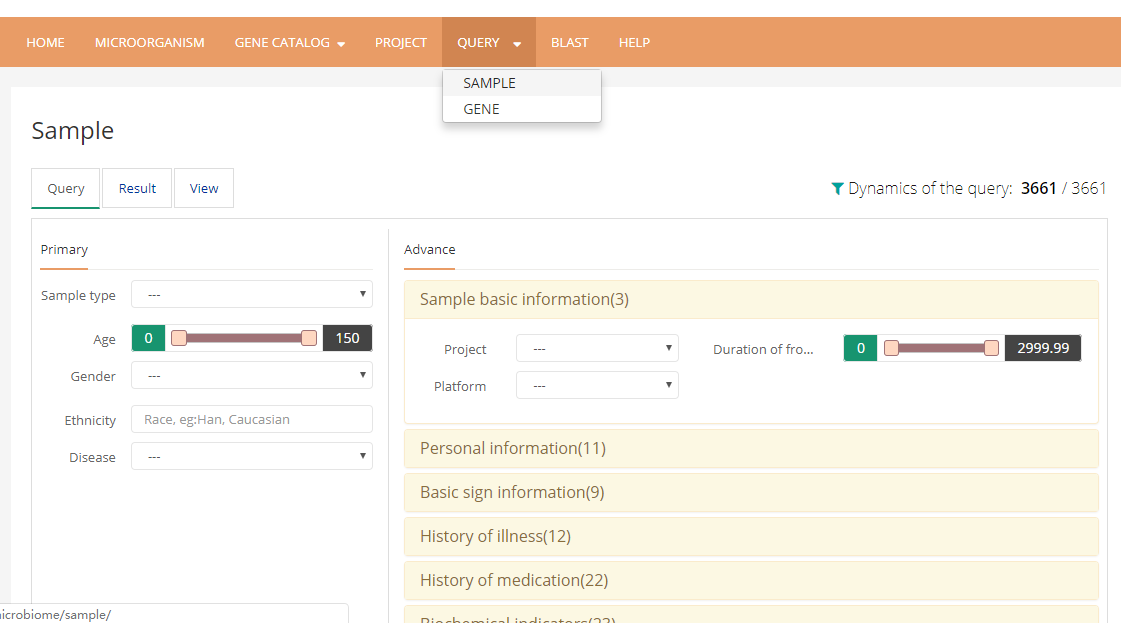
5 In the query page, user can choose interested samples by narrowing down parameters on more than 170 phenotypes.
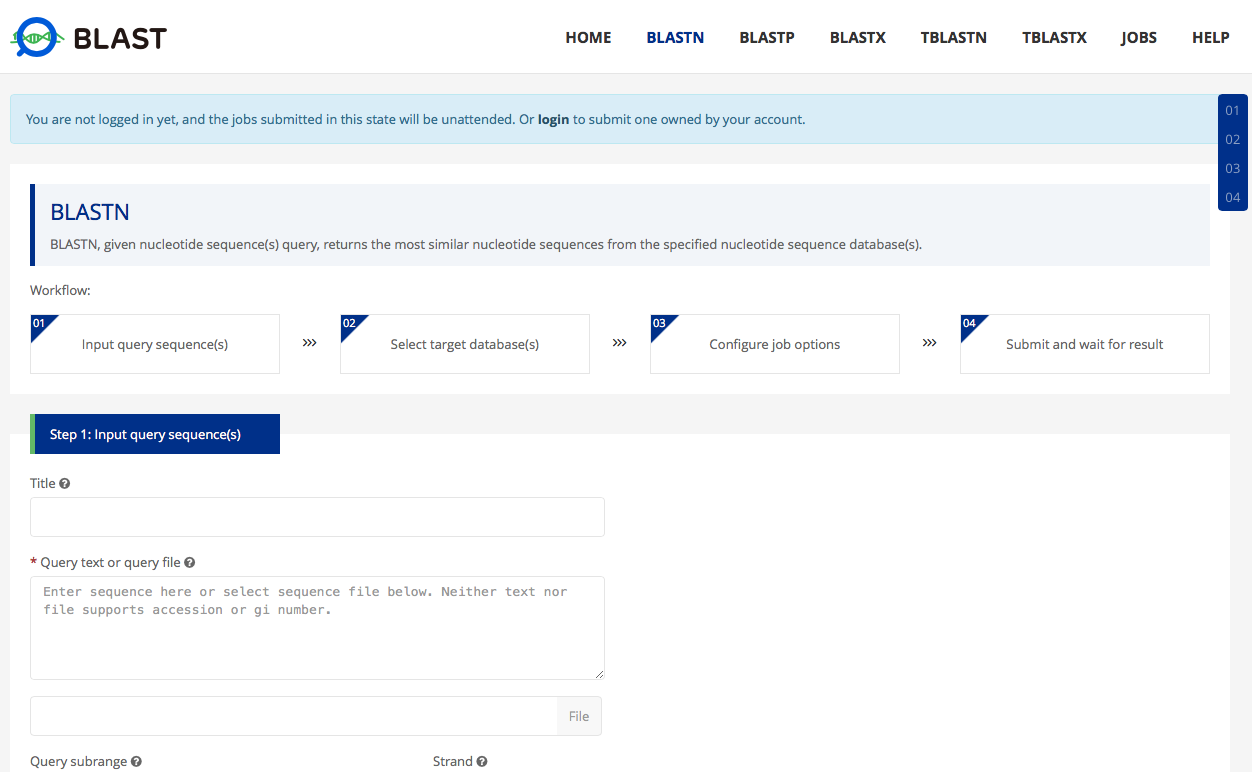
6 By clicking BLAST tag, user will be navigated to our BLAST website of CNGBdb. User can choose HMD projects from ‘project’ list on the right, and then user can paste their own nucleotide sequence or upload a file contains their sequence, to align against microorganism genomes in HMD or against microorganism gene sequence in our gene catalog. Hit records will be listed as result.
7 Citations:Please credit the Microbiome Database(MDB)if you use it in your work with the following link: db.cngb.org/microbiome